Blunt-end cloning: Ultimate guide
Introduction
Blunt-end cloning is a simple method to directly clone blunt-ended DNA fragments into a linearized plasmid with blunt ends.
Let us first understand what blunt-ends are. Blunt ends are the ends of a double-stranded DNA where nucleotides are perfectly paired (Fig 1). Commonly, the blunt-ends can be generated by the following means:
- Amplification of DNA using proofreading DNA polymerases.
- Digestion of DNA using blunt-end-generating restriction enzymes (REs).
- Modifying non-blunt ends into blunt-ends (End repair).
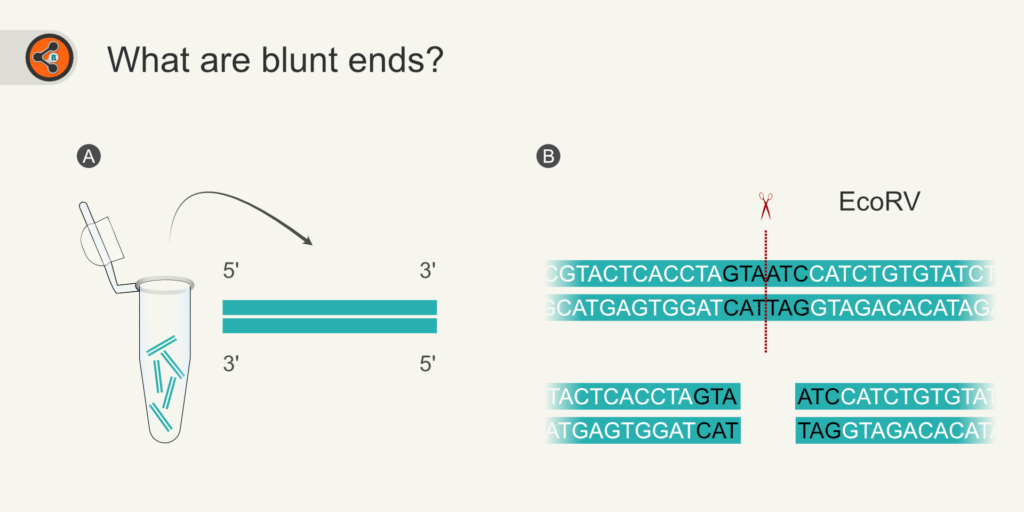
Figure 1: Illustration showing two major ways of generating blunt-ends. 1.1 Blunt-end generation by PCR amplification with proofreading DNA polymerase. 1.2 Restriction digestion of DNA using blunt-end generating enzymes (e.g., EcoRV).
As stated above, blunt-end cloning is the process of fusing insert and vector that are blunt-ended. Fig 2 represents the overview of blunt-end cloning. Incubating Blunt-end vector (Fig 2.2) with blunt-end insert and Ligase (DNA glue) results in a recombinant plasmid.
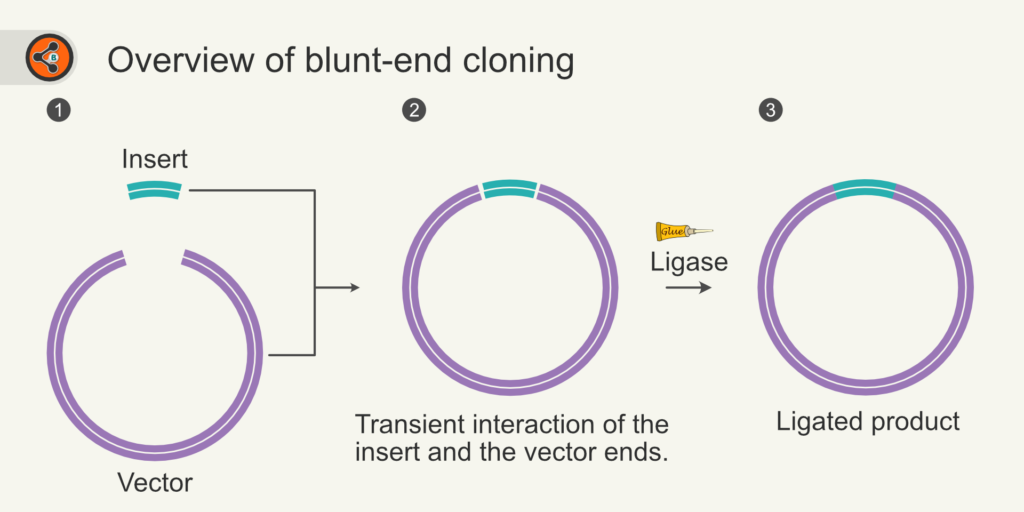
Figure 2: An overview of blunt-end cloning. 2.1 blunt-ended vector and insert. 2.2 Transient association of insert and vector. 2.3 Ligation of insert and vector resulted in a covalently closed plasmid.
The blunt-end ligation is a non-directional cloning method, i.e., the insert can ligate to the vector in two possible directions. Some workflows need the insert to be in one particular direction, e.g., protein expression. Usually, the desired directional clone can be identified using a PCR (using a combination of insert and the vector-specific primers). On the other hand, a variant of blunt-end cloning called the “directional blunt-end cloning” solves the directionality issue (discussed at the end of the article). In addition to this, a new ligase-free method utilizes topoisomerase to ligate the fragments, and it is named TOPO Blunt-end Cloning.
Principle
In blunt-end cloning, both the vector and the insert contain blunt-ends. During the transient association of the ends of the vector and the insert, the DNA ligase enzyme seals the joints. The T4 DNA ligase utilizes ATP to make a phosphodiester bond between the 3’ hydroxyl group of one DNA strand and the 5’ phosphate group of another DNA strand.
A detailed mechanism of ligation is discussed here.
Advantages
- Blunt-end cloning involves simple design of the primers (without any extra bases at the 5’ end of the primer).
- Blunt-end cloning has an advantage over other cloning methods (TA-cloning or traditional restriction enzyme-based sticky-end cloning). It does not require a complementary sequence between the insert and the. This property makes it a truly universal cloning method.
- It is used for sub-cloning, sequencing, building libraries of DNA molecules, and expressing coding and non-coding RNA.
Limitations
- Non-directional cloning generates only 50% of inserts with the proper orientation.
- In blunt-end cloning, both inserts and vectors do not have complementary 3’ or 5’ overhangs. This results in a little chance for a stable association between the insert and the vector, resulting in a lower recombination efficiency (10-100x) than sticky-end cloning.
- Blunt-end cloning is prone to vector self-ligation. Additional steps are needed to avoid self-ligation (discussed below).
- In certain instances where the insert concentration is high, a ligated vector can have more than one insert.
- It is challenging as the probability of generating a clone with multiple inserts with the right orientation is low. This also increases screening efforts to get the clone with the desired orientation.
Procedure – an overview
Blunt-end cloning involves the following steps.
- Preparation of Insert
- Preparation of Vector
- Removal of the terminal phosphate of the linearized vector (phosphatase reaction)
- Ligation
- Transformation
- Selection of transformants and other downstream processes.
Let us see all the steps in detail.
Preparation of the Insert
An Insert is any desired fragment of DNA that you want to clone into a vector/plasmid. It can be sourced from the genomic DNA, other plasmids, or any other source. Before the ligation reaction, the desired DNA fragment needs to be separated from its source as follows (Fig 3).
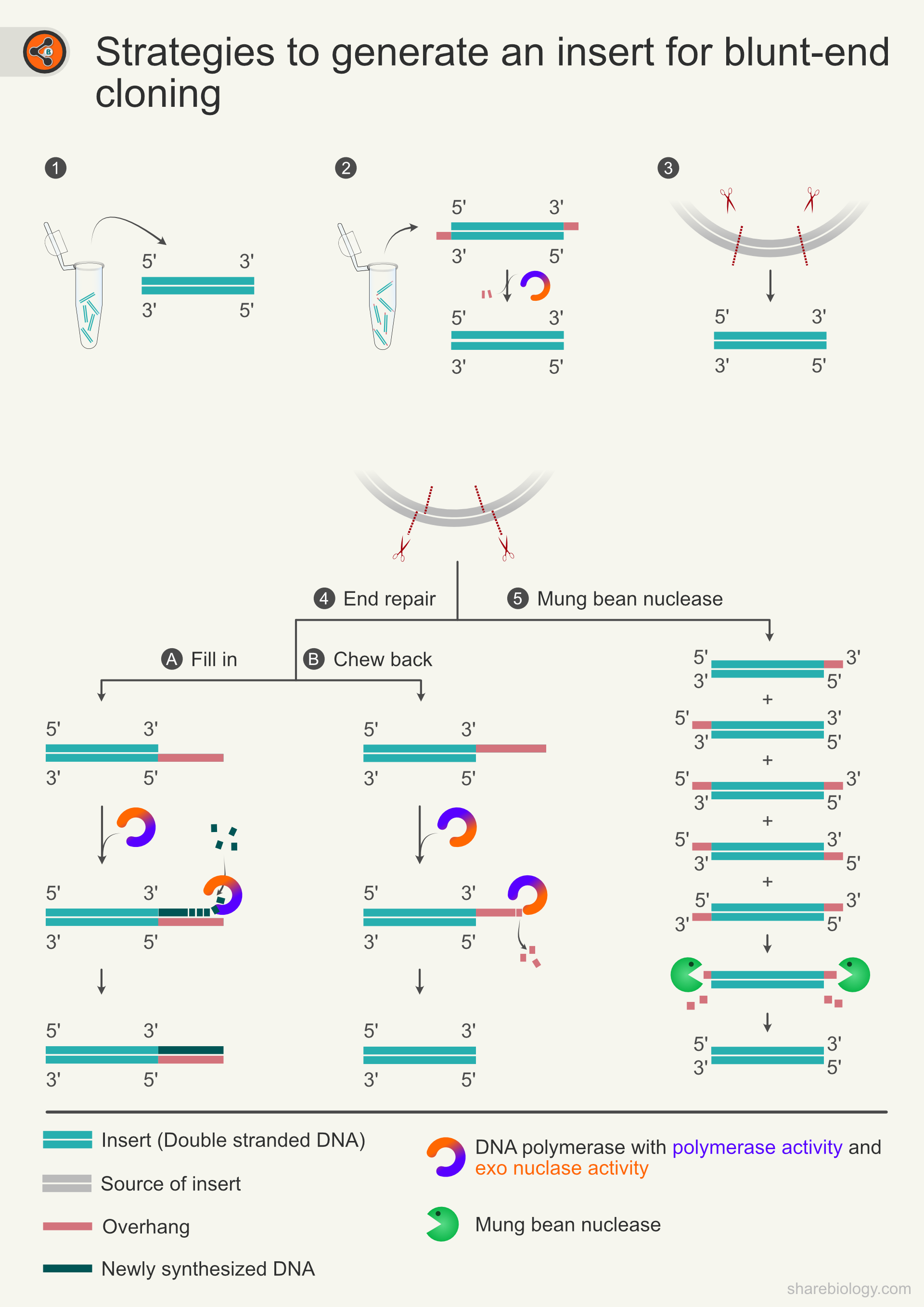
Figure 3: Strategies to generate an insert for blunt-end cloning. 3.1 One of the common ways to generate a blunt-ended insert is to amplify the desired piece of the template DNA using a proofreading polymerase. 3.2 Amplicons generated by a non-proofreading polymerase have overhangs (extra nucleotides on one strand of dsDNA) that need to be removed by PCR polishing. 3.3 Generation of blunt-end insert via restriction digestion with blunt-end generating enzymes. 3.4 If sticky-ends are generated in the workflow, they can be converted to blunt-ends by end repair (Fill in 3.4A and chew back 3.4B). 3.5 Alternative to end repair, mung bean nuclease generates blunt-ends via digestion of 3’ and 5’ overhangs.
- PCR amplification using proofreading DNA polymerase: Amplicons synthesized by the proofreading polymerase (3’->5’ exonuclease activity) will have blunt ends (Fig 3.1). Pfu, Phusion, KAPA HiFi, Q5 DNA pol, and Prime star GXL are a few examples of proofreading DNA polymerases. These amplicons can be used for ligation without the need for further modifications.
- PCR amplification using non-proofreading DNA polymerase: If the PCR product is amplified using non-proofreading polymerase (Example Taq DNA polymerase), amplicons will likely have a 3’ ‘A’ extra nucleotide. The presence of the extra ‘A’ prevents blunt-end ligation. The PCR product can be incubated for some time with the proofreading enzyme (PCR Polishing) (Fig 2) to remove the ‘A’ overhang.
Primer design
Blunt-end cloning primers do not require any special sequences or modifications. A detailed demonstration of primer designing is mentioned here.
- Using Blunt-end-generating restriction enzymes: Since the digestion products of blunt-end generating enzymes have blunt ends, they can be used for ligation (Fig 3.3).
- Use of sticky end generating restriction enzymes: Before ligation, sticky ends must be converted to blunt ends by end repair (Fig 3.4). During end repair, DNA polymerases such as T4 DNA polymerase or Klenow fragment are used for filling (Fig 4A) 5’ overhangs with polymerase activity and chew back (Fig 3.4B) 3’ overhangs with 3-5’ exonuclease activity. Whether the enzyme fills or chews back depends upon the type of overhang (5’/3’). These enzymes are preferred over proofreading polymerases as they have 5’à3’ exonuclease activity, which may lead to loss of information.
- Mung bean nuclease: Incubation with mung bean nuclease removes the sticky-ends of both 3′ and 5′ overhangs and turns the ends into blunt ends (Fig 3.5). This method is not recommended when the DNA sequences are essential, as the digested overhangs may have needed information.
If the DNA fragments are generated using sonication or shearing, the same filling or chew-back methods can generate blunt-end inserts.
Blunt-end vector preparation
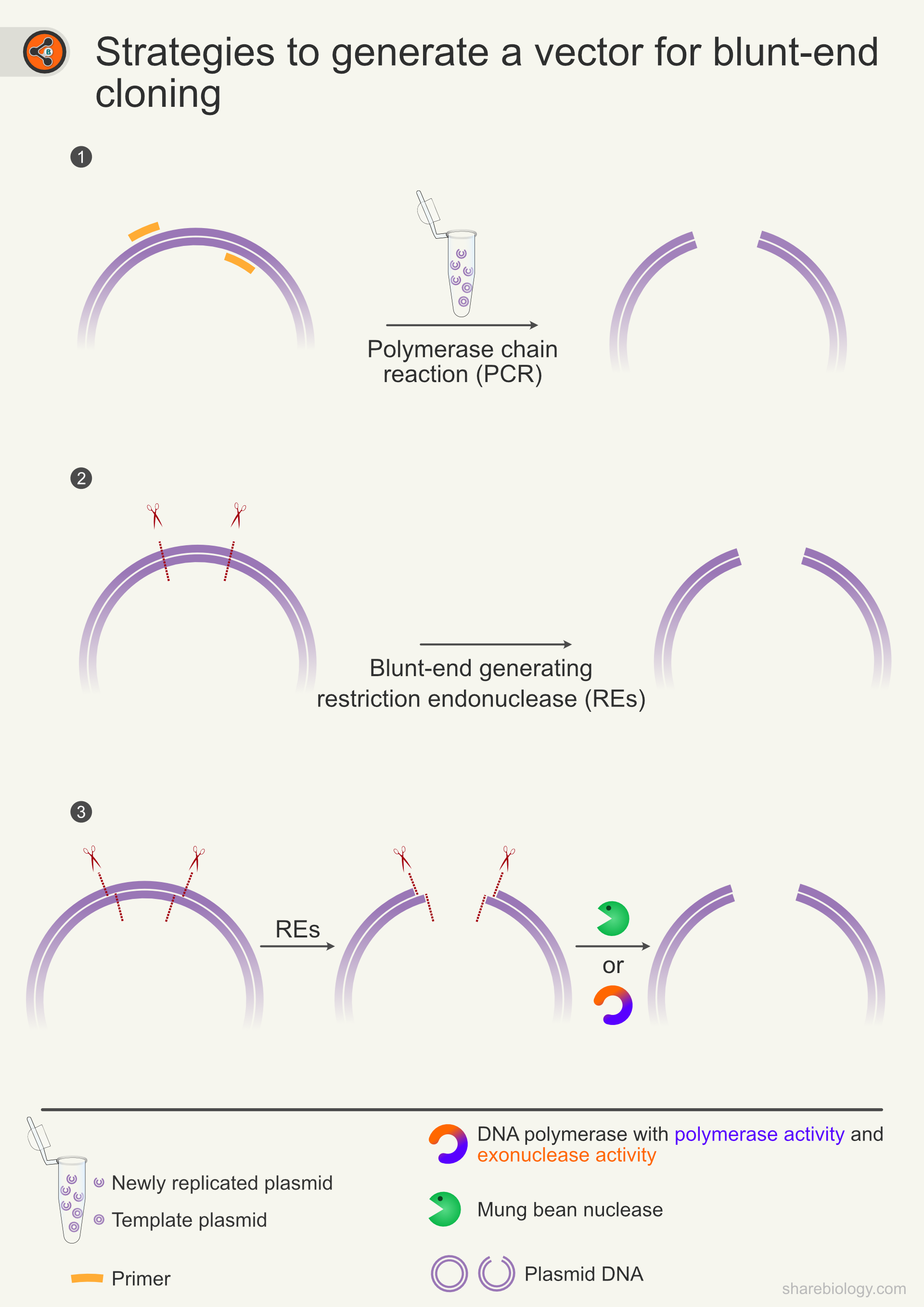
Figure 4: Strategies to generate a plasmid for blunt-end cloning. 4.1 One of the common ways to generate a blunt-ended vector is to amplify the desired piece of the template DNA using a proofreading polymerase. 4.2 Amplicons generated by a non-proofreading polymerase have overhangs (extra nucleotides on one strand of dsDNA) that need to be removed by PCR polishing. 4.3 If the vector has sticky-ends, they can be converted to blunt-ends either by end repair or mung bean nuclease.
Similar to the insert, the vector also can be prepared by different methods. The method of choice depends on whether the vector has blunt-end restriction sites or not.
- Amplification of the entire vector backbone using a proofreading polymerase (Fig 4.1).
- The circular vector can be linearized using restriction endonucleases that produce blunt ends (Fig 4.2).
- If the vector is linearized using restriction enzymes that generate cohesive ends, a filling or a chew-back mechanism can be used to turn them into blunt-ends (Fig 4.3).
- Use commercially available blunt-end vectors (Linearized vectors).
Phosphatase reaction of the vector before ligation
All the PCR amplified fragments (except when dephosphorylated primers are used), and restriction-digested fragments contain a 5’ terminal phosphate. The presence of terminal phosphate creates a significant problem in blunt-end ligation, i.e., self-ligation of the vector backbone. Self-ligation increases the background, i.e., transformed E.coli colonies with just the vector without the insert in it. This problem can be minimized by reducing the self-ligation possibility by removing phosphates present at the 5’ termini in the vector. The phosphatase treatment will effectively reduce the background of “empty” clones by >95%. Phosphatases are enzymes that do this job. Shrimp Alkaline Phosphatase (rSAP), Calf Intestinal phosphatase (CIP), and Antarctic Phosphatase (AnP) are widely used in labs. More details about the phosphatase reaction can be found in the protocol.
However, one should be cautious when using phosphatase reactions. It should be noted that for ligation to work, the ligase needs a phosphate group. So, a phosphate group must be present either on the insert termini or on the vector termini, and the lack of phosphate on both results in ligation failure. If the phosphate is not present at the 5′ end of either vector or insert, it can be added using T4 polynucleotide kinase. Usually, vectors are subjected to dephosphorylation reactions, as the self-ligated vector can propagate in the host, whereas the self-ligated insert cannot.
Ligation reaction
After purification of the fragments from previous reactions, the insert and the plasmid are set for ligation. The ratio of insert to vector is critical for optimal output. In general, a 3:1 molar ratio of insert to vector is preferred (molar ratio calculator).
The transient association of vector and insert ends (5’ phosphate group and 3’ OH group of double-stranded DNA) will be ligated by the ligase. Only the T4 DNA ligase enzymes can ligate the blunt-ends. Taq-ligase and several other ligases do not ligate blunt-ends or at the least need very stringent conditions.
It should be noted that only one phosphodiester bond is allowed to be formed at each junction due to the dephosphorylation of the vector. So, one strand attaches to the vector at each junction while the other is nicked. Overall, only two phosphodiester bonds hold the vector and insert. The nicks will be repaired in the cell after transformation.
How to increase the efficacy of blunt-end cloning?
- Extended incubation periods, in general, increase the ligation efficiency.
- PEG (polyethylene glycol) may increase efficiency by increasing molecular crowding.
- Blunt-end cloning is less efficient as the vector and the insert do not have any complementary sequence. Liu ZG in 1992 showed increased efficiency of blunt-end cloning by incubating the ligation mixture in the presence of blunt-end restriction enzyme (Sma1). The idea is that the self-ligation of vector generates restriction enzyme recognition sequence, whereas the ligation of vector-ends with insert does not. The self-ligated vectors get digested again until they get ligated with insert. This strategy works only if the vector is generated using a blunt-end restriction enzyme. It is to be noted that, upon ligation, the afore-mentioned restriction site should not be created, and the insert should be devoid of the same.
- Similar efficiency can be achieved by treating the vector-ends with the Phosphatase enzyme, which inhibits the vector self-ligation (see phosphatase reaction).
- Bercovich, in 1992 used equimolar concentration of vector:insert to increase the efficiency of cloning. He used a total DNA concentration of <100 μg/mL.
- Incubation of PCR amplified products using T4 polynucleotide kinase ensures the presence of 5’ terminal phosphate.
Directional blunt-end cloning
Directional blunt-end cloning is facilitated by amplifying both the insert and the vector using a set of primers consisting of a phosphorylated primer and a dephosphorylated primer. During the ligation reaction, the insert can be ligated only in one direction as the reverse orientation does not have phosphates either on the vector end or the insert end for the ligation to happen.
References
- An efficient method for blunt-end ligation of PCR products.
- Effect of DNA concentration on recombinant plasmid recovery after blunt-end ligation.
- Protocols for cloning and analysis of blunt-ended PCR-generated DNA fragments.
- Molecular Cloning: A Laboratory Manual. 4th ed.
- NEB (New England Biolabs).
- Bitesizebio.
- arcticzymes.com.
- Integrated DNA technologies
