Blunt-end TOPO Cloning

Introduction
Blunt end cloning is a simple method to directly clone PCR amplified products and fragments isolated from restriction digestion of plasmid or genomic DNA into a blunt-ended vector. Both vector and insert should contain blunt ends for a successful cloning reaction.
TOPO blunt cloning is restriction enzyme-free and ligation-free cloning. Tradition blunt-end cloning method requires ligation step to seal the joints of the vector with insert. The ligation step is skipped in TOPO blunt cloning using the topoisomerase 1 enzyme. In 1987 Shuman and Moss identified the DNA topoisomerase 1 from the vaccinia virus. The topoisomerases relax the super-coiled DNA by unwinding or winding duplex DNA by cutting and rejoining during replication and transcription.
The topoisomerase 1 from the vaccinia virus recognizes the pentapyrimidine element 5′-C/TCCTT-3′ sequence and cuts the DNA (Shuman et al. 1991, 1992). The ability of the topoisomerase enzyme to identify specific DNA sequence, cutting within the recognition sequence, and rejoining was utilized in TOPO cloning.
Only the proofreading DNA polymerase (3′ to 5′ exonuclease activity) like Pfu should be used to generate Blunt end PCR products. The non-proofreading polymerase like Taq pol does not generate blunt end products.
Principle
DNA topoisomerase enzyme has sequence-specific DNA cleavage and ligation activity. The addition of Topoisomerase enzyme to ends of the vector can be utilized to ligate these ends with insert. The topoisomerase 1 from the Vaccinia virus recognizes 5’-C/TCCTT-3′ sequence and cuts it. In the cleavage reaction, topoisomerase is attached to the 3’end by forming a covalent bond between cleaved 3′ DNA strands to topoisomerase’s tyrosine residue (Tyr274). The energy stored in this phospho-tyrosyl bond between the DNA and enzyme utilized to religate the ends that originally cleaved or relegate to a heterologous acceptor DNA (Insert).
The enzyme gets released from vector ends after rejoining the ends. In 1994 Shuman used this topoisomerase 1 from the vaccine virus enzyme to make recombinant plasmid.
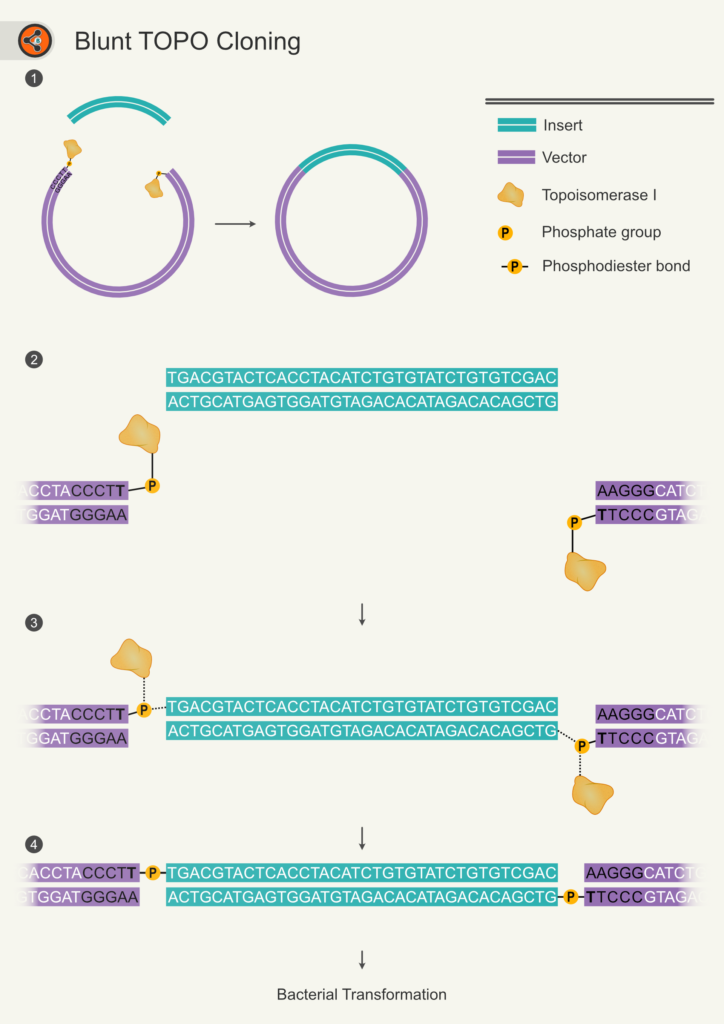
Figure 1. Schematic diagram representing steps in Blunt-end TOPO cloning. 1.1 An overview of Blunt-end TOPO cloning. Insert (teal) is ligated with vector (violet) resulting in recombinant vector (final product). 1.2-1.4 Shows the steps involved in the ligation during topo cloning. 1.2 Shows the insert with blunt-ends and the linearized blunt-end vector with topoisomerase attached to 3′ T (shown in bold) of the CCCTT sequence present at both ends. 1.3 The 5′ OH of inset grabs phosphate group via nucleophilic substitution and replaces topoisomerase I with itself. 1.4 After successful TOPO reaction, insert attached with the vector backbone with two phosphodiester bonds in total. This arrangement leaves nicks that would be sealed in the bacteria after transformation.
Advantages
- It is an easy-to-use, simple cloning technique. A better alternative to blunt-end cloning.
- Restriction-free and ligation-free cloning.
- The simple design of primers without any additional bases at 5′ end.
Limitations
- Non-directional cloning generates inserts only 50% in the proper orientation.
- Dependency on the vendors for TOPO blunt vectors; it will be challenging to create vectors in the lab.
Procedure – an overview
Blunt-end TOPO cloning involves the following steps.
- Preparation of Insert
- Selection of vector
- TOPO cloning reaction
- Transformation
- Selection of transformants and other downstream processes.
Let us see the first three steps.
Preparation of Insert
The detailed description of blunt end insert preparation is discussed here.
Primer design
Blunt-end TOPO cloning primers do not require any special sequences or modifications. A detailed demonstration of primer designing is mentioned here.
Vector
The vector needs to be purchased from the vendor (Blunt end cloning master mix). Vectors are available with different feature sets that aids in downstream processes. Please choose a suitable vector as per your needs.
TOPO Reaction
Mix the blunt-ended insert with TOPO reaction mixture, which contains PCR blunt II topo vector. Incubate the reaction sample at 25°C for 15 min and proceed with transformation.
Transformation
Detailed protocol for transformation is discussed here.
