TA cloning: Ultimate guide
Introduction
TA cloning is a simple method to clone any desirable fragment with an extra A (Adenine nucleotide) overhang into any linearized vector with T (Thymidine nucleotide) overhang.
The T and A overhangs are of paramount importance in TA cloning. Let us learn how they are added to the DNA.
DNA polymerases are enzymes that synthesize new DNA strands using a template strand. However, some polymerases add nucleotides without using any template (non-template-dependent polymerization/ non-template addition of nucleotides).
The non-template-dependent addition of one or more nucleotides at the 3′ end of PCR-amplified products was first observed by Clark et al. The work of Hu et al. states that the type of the nucleotides added depends on the DNA sequence and type of polymerase (more on this later!). Like DNA polymerases, the terminal transferases are enzymes that add one or more nucleotides to the 3′ end. The addition of extra nucleotides to the blunt ends is called Tailing.
T.A. Halton developed the first successful TA cloning method in 1990. With this method, Halton bypassed the use of restriction enzymes. In conventional cloning, PCR amplified fragments were digested using restriction enzymes to generate compatible cohesive ends with the vectors.
Halton generated the T overhang-containing pBluescript vector in his lab using the ddTTP (dideoxythymidine triphosphate) method. He first digested the vector using EcoRV, a blunt-end restriction enzyme, to generate a blunt-end vector and performed the tailing reaction using terminal transferase. The absence of the 2’OH group of ddTTP makes sure that only one thymidine is added at the 3′ end by the Terminal transferase enzyme. He mixed this T-overhang containing vector with an A-overhang containing PCR fragments (Plasmid cDNA library of petunia petals) amplified by Taq polymerase. 80% of the transformants are positive for the insert.
In the subsequent year, i.e., 1991, Marchuk et al. simplified the process of addition of T-overhang to the plasmid vector by using Taq DNA polymerase and dTTP. Unlike terminal transferase, the Taq polymerase adds only one nucleotide (preferably adenine). Providing only dTTP in the tailing reaction allows T-overhang to get added instead of A. He first digested and used the pBluescript vector using EcoRV, a blunt end restriction enzyme, to generate a blunt-end vector. He then incubated this vector in a PCR reaction supplied only with dTTP to generate a T-overhang.
The traditional TA cloning method is ligase-dependent cloning. TOPO TA cloning is a better version of TA cloning that is ligase-independent with a simple workflow.
Principle
In the TA cloning method, the Adenine nucleotide at the 3′ end of the insert will pair with the vector Thymidine nucleotide. During the stable association of vector and insert ends using AT base-pairing, the DNA ligase enzyme seals the joints. The DNA ligase utilizes the ATP or NAD+ to make a Phosphodiester bond between the 3’OH group of adenine and 5′ phosphate group of another nucleotide. Two phosphodiester bonds are formed to join the DNA fragments (ds DNA) at each junction.
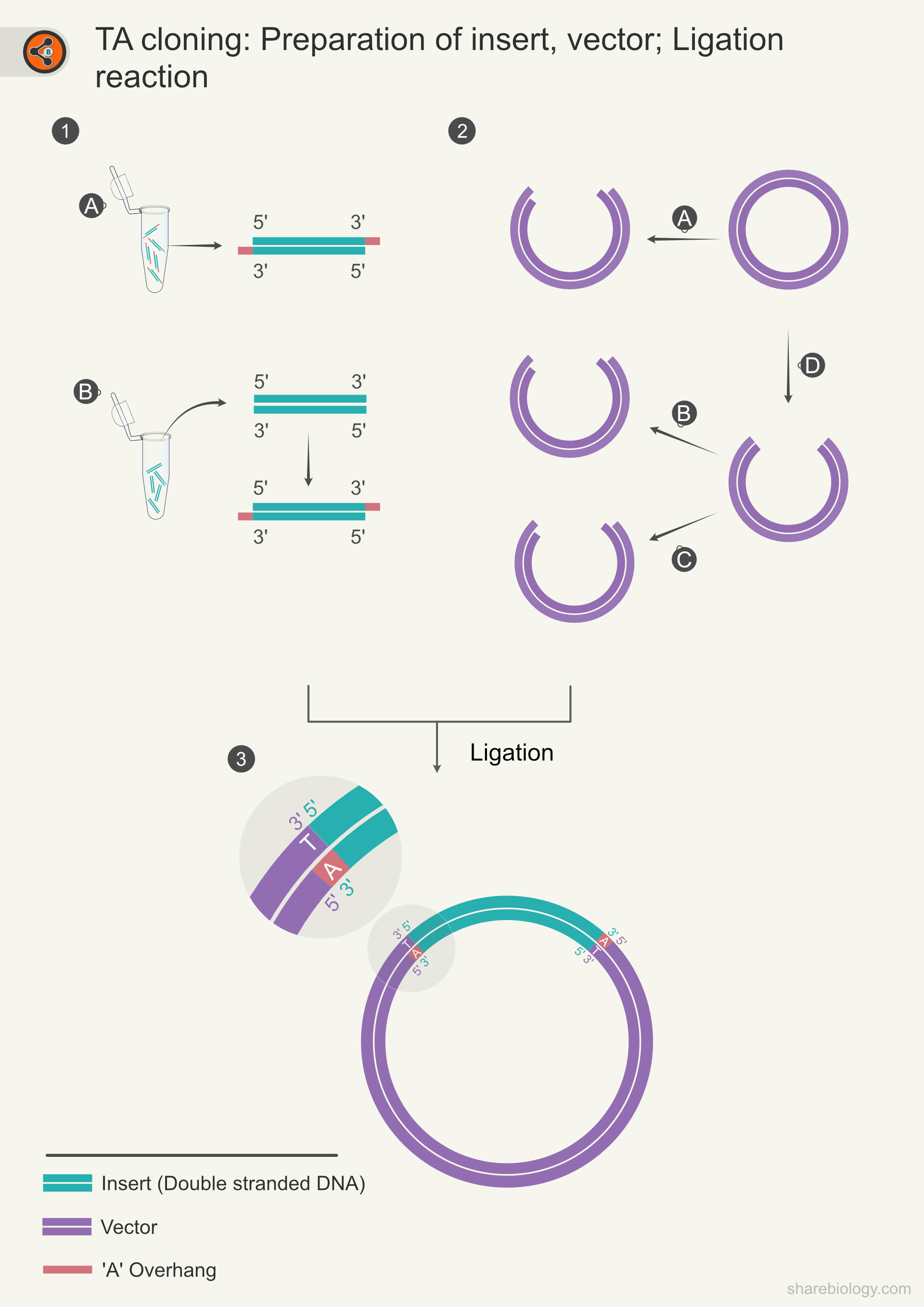
Figure 1: Illustration representing the steps in TA cloning. 1.1 Represents the preparation of insert. 1.1A Amplification of the insert with Taq DNA polymerase results in an A-overhang. 1.1B Adding an A-overhang to the blunt-end insert using Taq DNA polymerase and dATP in a tailing reaction. 1.2 represents vector preparation. 1.2A vector is linearized using restriction enzymes that generate T overhangs. 1.2D In some workflows, it is not possible to utilize restriction enzymes to generate T-overhangs. So the vectors are first linearized with blunt-end generating enzymes, then these vectors are further processed to have T-overhangs. 1.2B Addition of T-overhang using Taq DNA polymerase and dTTP. 1.2C Addition of T-overhangs with terminal transferase and ddTTP combination. 1.3 The ligation reaction creates the recombinant vector.
Advantages
- It is easy to use and is a simple cloning technique generally used for subcloning.
- TA cloning does not require extensive restriction enzymes compared with conventional, golden gateway cloning.
- The simple design of primers (without any extra bases at the 5′ end of primer).
- The cloning efficiency of TA cloning is more than that of blunt-end cloning.
- Much simpler, faster than traditional cohesive end cloning because Vectors and PCR amplified products do not require extensive restriction digestion steps.
- This method does not require any phosphatase reaction as the vector is not blunt-ended.
Limitations
- Non-directional cloning generates only 50% of inserts in the proper orientation.
- TA cloning kits are expensive because vectors with T overhang need to be purchased. T- overhangs can be generated in the lab (Procedure given below) but are usually bought from the vendors.
- TA cloning is ligation-dependent cloning, when compared with TOPO TA cloning, requires an additional step.
Procedure – an overview
TA cloning involves the following steps.
- Amplification or preparation of insert
- Selection or generation of T overhang containing the vector
- Ligation
- Transformation
- Selection of transformants and other downstream processes.
Let us see the first three steps in detail.
Amplification or preparation of insert
The insert for TA cloning can be a DNA fragment that comes from PCR amplification or the digestion of plasmids, genomic DNA, or other sources. Whatever the source of the insert is, it must contain an A-overhang. If the insert is blunt-ended, go to step two (B) and add A-overhang.
Following is the procedure to generate insert using PCR amplification.
- If the PCR product is amplified using non-proofreading polymerase (e.g., Taq polymerase), they generate PCR products with a 3′ A-overhang (fig 1.1A). The downside is that they may cause mutations in the amplicons.
- The PCR products are amplified using proofreading polymerase (g., Pfu polymerase) with 3′-5′ exonuclease activity, generating blunt-end products. Incubation of this purified blunt-end PCR product with regular Taq polymerase in the presence of dATP causes the addition of A-overhang (Fig 1.1B).
Primer design
TA cloning primers (not to be confused with insert) do not require any special sequences or modifications. A detailed demonstration of primer designing is mentioned here.
Selection or generation of T overhang containing the vector
There are three ways to generate vectors for TA cloning
A. Digestion of circular vectors using the following restriction enzymes generates T-overhangs; the linearized vector can be directly used for cloning. (HphI (Mead et al., 1991), AhdI (Jeung et al., 2002), and Bfu I (Janner et al., 2013; Wang et al., 2013a) (Fig 1.2A).
B. Incubation of blunt-end vector (Fig 1.2D) with Taq DNA polymerase and addition of only dTTP result in the addition of thymidine at 3′ ends (Fig 1.2B). This protocol should work because given an equimolar amount of dNTPs in a PCR reaction, Taq DNA polymerase I will add an extra (template-independent) A. If the enzyme is supplied with only dTTP, then an extra T will be added (Marchuk et al. 1991).
C. Incubation of the blunt-end vector with Terminal transferases enzyme in the presence of ddTTP (do not use dTTP, as it makes poly T-tail) results in the addition of single T at the 3′ end (Fig 1.2C). (Ref: A simple and efficient method for direct cloning of PCR products using ddT-tailed vectors. T A Holton and M W Graham).
Use commercially available vectors for TA cloning. Addgene provides an extensive list of TA cloning vectors. Some of the regularly used T vectors are provided at the end of this article.
Factors influencing non-template addition of 3’A
Hu et al. showed that the non-template-dependent addition of nucleotide at the 3′ end is affected by both DNA polymerase and the last nucleotides. The proofreading polymerases like pfu DNA polymerase and T4 DNA polymerase do not add extra nucleotides and generate blunt ends. The Taq DNA polymerase, Vent DNA polymerase, T7 DNA polymerase, DNA polymerase I from Escherichia coli, and Klenow fragment (exo-) add an extra nucleotide. The extra nucleotide is generally Adenine (Adenylation). The percentage of overall adenylation of PCR amplified products is dependent on the last nucleotides of the 3′ end.
According to this study, the presence of Cytosine or adenine promoted adenylation, and the presence of guanine promoted the addition of G and A. In contrast, the presence of thymine resulted in the replacement of thymine with adenine.
| 3’ end residue(s) | Effect on Adenylation ( +A ) |
|---|---|
| AAA or A | +A (low efficiency) |
| GGG or G | +G> +A>C |
| CCC or C | +A > +C |
| TTT or T | -T, +A |
However, Hu et al. study did not mention the effect on adenylation beyond the first nucleotide. L. Magnuson filled this gap in 1996. He analyzed the impact of the last three residues using ~90 combinations of the last three residues. Their results show that the adenylation is influenced not only by the last nucleotide (consistent with Hu et al.) but also by other adjacent nucleotides. They did observe that if Thymine is present at the terminal end, adenylation happened without the loss of Thymine as contrary to hu et al. The difference between both the studies may be because of Buffer composition (Magnesium ion concentration) and PCR conditions.
| 3’ end residue(s) | Percentage of Adenylation (+A) |
|---|---|
| AAA | 24 |
| GCA | 24 |
| CAA | 31 |
| GCA | 29 |
| TCA | 32 |
| CCC | 84 |
| AAT | 84 |
| AAC | 83 |
| CTT | 79 |
| ACC | 77 |
Ligation
After purification of the fragments from previous reactions, insert and plasmid DNA is set for ligation. The ratio of insert to vector is critical for the optimal output. In general, a 3:1 molar ratio of insert to vector is preferred.
In the ligation reaction mixture, transient association of insert and vector occurs. This association is relatively stable compared to that of blunt-end insert-vector association as T and A overhangs of insert-vector are held by hydrogen bonds. This transient complex becomes stable by the formation of covalent bonds between the 5′ phosphate group and 3′ OH group of insert and vector. Ligase mediates this reaction. Unlike blunt-end cloning, which requires T4 DNA ligase, TA cloning can be performed by using any ligase.
After the ligation, two phosphodiester bonds are formed at each junction (vector-insert). In blunt-end cloning, each vector-insert junction is held by one phosphodiester bond as the vector is dephosphorylated.
TA cloning vectors
There are 300+ T overhang containing vectors available for TA cloning (Addgene)
Some examples of Commercial TA vectors are listed below.
Vectors for subcloning
pMDTM (TaKaRa)
pMD20 and pMD19 (TaKaRa)
pGEM-T and pGEM-T Easy Vector systems (Promega)
pCRT and pCRZeroT (DOI: 10.1038/s41598-019-42868-6)
pCAMBIA1300 (DOI: 10.1104/pp.109.137125)
pGreen vectors (DOI: 10.1371/journal.pone.0059576)
pPCV (DOI: 10.1186/2193-1801-2-441)
pBS2ndd and pBS3ndd (DOI: 10.1007/s11274-018-2466-z)
pKILPCR (DOI: 10.2144/97235pf01)
Vectors for protein expression (expression vectors)
pTARGETTM Vector from Promega,
pDRIVE Vectors from InvivoGen,
pUCm-T Vector from Sangon
References
- Clark, J. M. (1988). Novel non-templated nucleotide addition reactions catalyzed by procaryotic and eucaryotic DNA polymerases. Nucleic acids research, 16(20), 9677-9686.
- Holton, T. A., & Graham, M. W. (1991). A simple and efficient method for direct cloning of PCR products using ddT-tailed vectors. Nucleic acids research, 19(5), 1156.
- Hu, G. (1993). DNA Polymerase-catalyzed addition of nontemplated extra nucleotides to the 3′ of a DNA fragment. DNA and cell biology, 12(8), 763-770.
- Liu, Q., Dang, H. J., Wu, Y. H., Li, M., Chen, Y. H., Niu, X. L., … & Luo, L. J. (2018). pXST, a novel vector for TA cloning and blunt-end cloning. BMC biotechnology, 18(1), 1-7.
- Marchuk, D., Drumm, M., Saulino, A., & Collins, F. S. (1991). Construction of T-vectors, a rapid and general system for direct cloning of unmodified PCR products. Nucleic acids research, 19(5), 1154.
- Magnuson, V. L., Ally, D. S., Nylund, S. J., Karanjawala, Z. E., Rayman, J. B., Knapp, J. I., … & Collins, F. S. (1996). Substrate nucleotide-determined non-templated addition of adenine by Taq DNA polymerase: implications for PCR-based genotyping and cloning. Biotechniques, 21(4), 700-709.
