This article is a summary of:
Podcast available for this article
This article presents the study of chloroplast development in the Arabidopsis cue8 mutant. An elegant genetic screen developed over 20 years ago in the Chory lab to study light-induced nuclear gene expression identified as ‘CAB gene under-expressed’ or ‘cue’ mutants. Among them, cue8 has a plastid development defect, and the mutant plants display virescence (slow greening) phenotype (Figure 1). Compared to more mature cells in the leaf tip, the newly emerging cells in cue8 leaf primordia and the cells at the base of the developing leaves show reduced chlorophyll.
Virescence and chloroplasts
We tracked the slow greening phenotype (cue8) by examining the chloroplasts at different developmental stages. The mesophyll cells of cue8 cotyledons and emerging leaves are less filled with small and immature plastids, and this results in the emerging/young leaves being pale. The cells of young cue8 leaves also had relatively lower plastid proportion while only the mature leaves showed chloroplast occupancy almost similar to that of the wildtype. The greening gradient of cue8 suggests a delay in chloroplast development, chloroplasts fill the available cellular space gradually and this slow the overall plant growth.
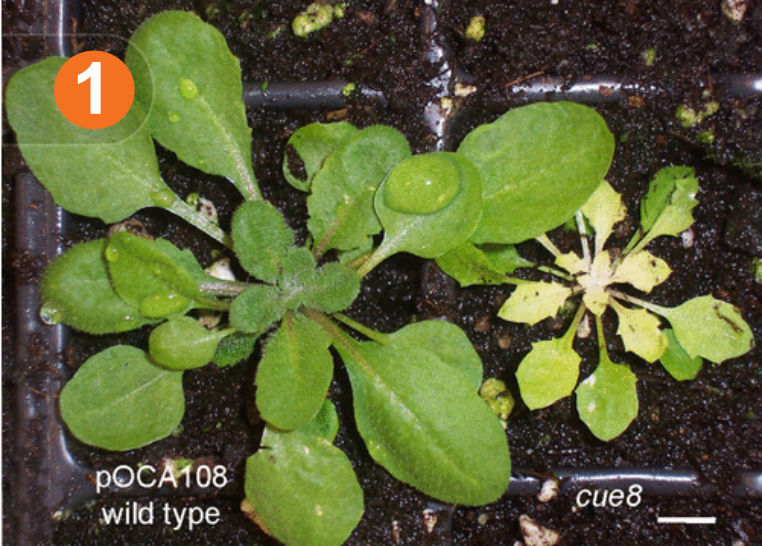 Figure 1. The phenotype of the Arabidopsis cue8 mutant and its wildtype
Figure 1. The phenotype of the Arabidopsis cue8 mutant and its wildtype
Retro-anterograde compensatory mechanism (gene expression, plastid genome copy number)
To build a fully functional chloroplast, cells need to produce about 3000 proteins. A majority of these proteins are encoded by the nuclear genome and are imported from the cytosol into the chloroplasts. The chloroplast genome codes for only about 80 proteins. Transcription of the chloroplast genome involves two types of RNA polymerases: a single-subunit Nucleus-Encoded Polymerase or NEP, similar to the one in mitochondria; and a multi-subunit bacterial-type Plastid-Encoded Polymerase or PEP. Like bacterial polymerase, the PEP enzyme requires the assistance of a sigma factor. There are 6 sigma factors in Arabidopsis, and they all are encoded by the nucleus. Studies show that NEP activity is high during the early plastid development; more importantly, NEP transcribes the genes for PEP subunits after entering the chloroplast. Although both polymerases are active in the chloroplasts, PEP seems to be predominantly responsible for the expression of photosynthesis-related genes, whereas NEP takes charge of the expression of early housekeeping genes.
As the phenotype suggests, cue8 shows a reduced expression of nuclear genes specifically coding for photosynthesis-related proteins and their corresponding Golden-2-like transcriptional regulators. Accordingly, the expression of photosynthesis-related genes in the plastids, transcribed by PEP, is also reduced. To our surprise, the expression of chloroplast genes transcribed by NEP increased in the defective plastids of cue8; genes which include those for the PEP subunits. It is known that the changes in plastid gene expression in seedlings caused either by genetic defects or by the treatment with lincomycin, a translation inhibitor, also show such a phenotype. We wanted to find out why there is less PEP-driven transcription despite there being many transcripts for PEP subunits, so we looked into the expression of sigma factors and found that the transcript levels of SIG2 and SIG5 were indeed reduced in cue8. Interestingly, the expression of NEP itself was enhanced 2-4-fold in cue8. Furthermore, when the number of chloroplast genomes per haploid nuclear genome was measured, the number of chloroplast DNA copies per cell in cue8 was found similar to that of WT chloroplasts. As a result, the cpDNA was observed densely packed in the immature chloroplasts of cue8, whereas it is scattered in WT chloroplasts. This evidence indicated that the increased expression of NEP-driven transcripts in cue8 is due to the increased expression of NEPs themselves and the maintenance of the number of chloroplast DNA copies in spite of the delayed chloroplast development relative to the WT.
We interpret this, the molecular phenotype of enhanced expression of chloroplast housekeeping genes by enhanced expression of NEPs and the maintenance of the DNA replication of defective plastids, as part of a retro-anterograde, compensatory mechanism that actively keeps the cue8 plastids in their juvenile stage.
Chloroplast to nucleus communication in cue8
The development of a proplastid into a mature plastid is under the control of the nucleus-encoded proteins and this is termed as “anterograde control”. Defects in plastids are communicated back to the cell’s nucleus through a process called “retrograde signalling”.
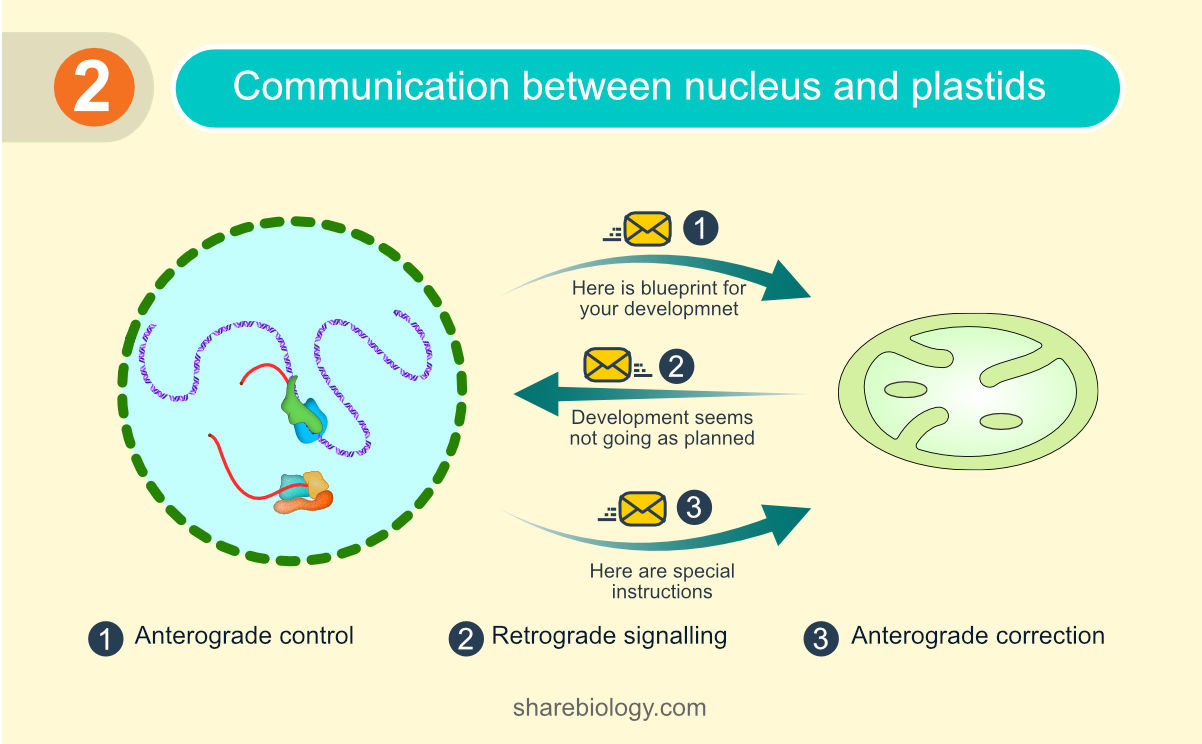
Figure 2. Schematic representation of communication between the nucleus and the chloroplast.
This plastid-to-nucleus backward communication shuts down the synthesis of nuclear-encoded photosynthetic proteins, which are either harmful (producing toxic compounds like reactive oxygen species or nitrite) or potentially produced in greater amounts than required because of plastid defects. Studies on retrograde signalling revealed a nucleus-encoded, chloroplast-localised protein, GUN1, as a key regulator and that interacts with other proteins involved in tetrapyrrole biosynthesis, plastid gene expression, and protein import in the chloroplast. It has been reported that plastid defects lead to changes in mRNA editing in several chloroplast transcripts. Recent studies also found that GUN1 interacts with, and alters the activity of a protein involved in chloroplast mRNA editing.
We asked whether cue8 defect is retrograde-signalled with the help of GUN1; for this, we generated a cue8 gun1 double mutant which turned out to be non-viable, lethal. This was at first a surprise to us, and it proves that retrograde signalling causes more than a small benefit of the “economy of resources”. It is essential for plant survival if plastids are defective. We looked for the presence of the compensatory response in those mutants. This brought a second surprise: the double mutants showed even more reduction in transcription of the nuclear LHCB gene and its GLK regulators. However, the elevation of NEP transcripts remained present in these mutants. The level of expression of chloroplast genes transcribed by the NEPs was not elevated as it did in cue8, but remained at WT levels; that component of the response had failed. The question of why there is no increase in NEP-driven transcripts was explained by the failure to accumulate a sufficient number of plastid DNA template copies.
It appears some aspects of the compensatory response, such as elevated transcripts of the NEP and organellar DNA POL and the reduction of nucleus-encoded photosynthesis genes, are maintained in cue8 even without GUN1; but others, such as elevated NEP activity and the overall maintenance of nucleoids, do require GUN1. This impaired corrective response coincides with the complete failure of chloroplast development and eventual seedling lethality of the double mutant. mRNA editing is a known but still surprising phenomenon by which some single bases change in organelle transcripts (specific cytosines become uracils), altering the sequence of the protein encoded. We find the issue of changes in mRNA editing in the chloroplast extremely intriguing; and we looked at what impact loss of cue8, with or without GUN1, had on mRNA editing. The result was also intriguing: loss of cue8 altered the frequency of editing, triggering increased editing in a housekeeping gene, rpoC1 yet reduced in a photosynthetic gene, ndhD; and the changes in the frequency of editing were even more pronounced in the plastids of albino cue8 gun1 double mutants.
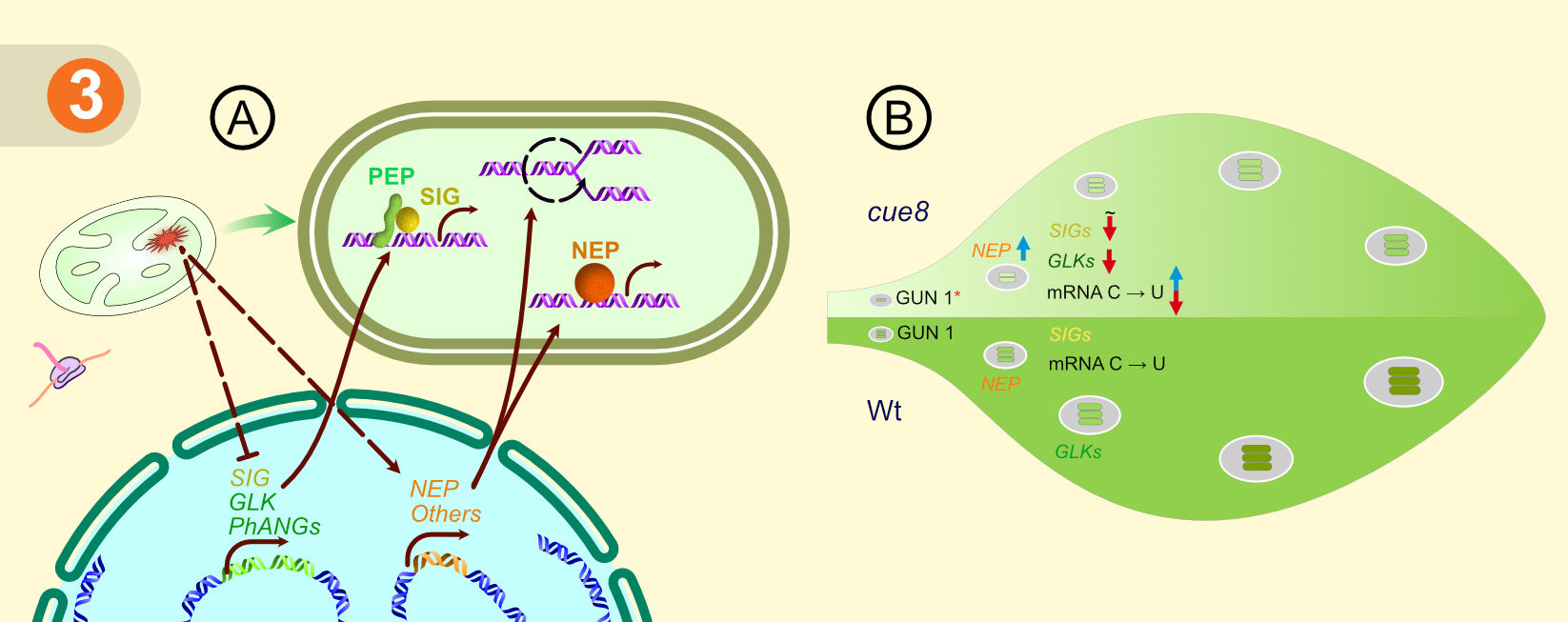
Figure 3. Summary of the cue8 molecular phenotype. (a) Plastid biogenesis defect in cue8 is communicated to the nucleus, and this results in reduced expression of nuclear genes associated with photosynthesis and sigma factor genes, and increase in the expression of NEPs and maintaining cpDNA copies. (b) The delayed greening phenotype of cue8 showing prolonged chloroplast development.
In summary, the response observed in cue8 resembles a prolonged juvenile state of early chloroplast biogenesis. The response follows retrograde signalling; it allows for anterograde correction manifested as a slow-greening phenotype; and it makes it possible, only if this happens, for relatively healthy chloroplasts to develop (Figure 3). Without this response, chloroplast development suffers catastrophic failure and the seedlings die. This response can almost be described as the operation of a “chloroplast development checkpoint”, and it requires GUN1 but only for a part of the response. GUN1 is not the complete story because some aspects of the response occur even without it.
*Images were reproduced or redrawn with permission.
References:
Loudya, N., Okunola, T., He, J., Jarvis, P. and López-Juez, E., 2020. Retrograde signalling in a virescent mutant triggers an anterograde delay of chloroplast biogenesis that requires GUN1 and is essential for survival. Philosophical Transactions of the Royal Society B, 375(1801), p.20190400. https://doi.org/10.1098/rstb.2019.0400
Naresh Loudya discusses nucleus-chloroplast signaling
[simple-author-box]